 Tutorials covering the estimation of expresson signatures of reference cell types (1/3), spatial mapping with cell2location (2/3) and the downstream analysis (3/3) can be found here: The model then decomposes spatially resolved multi-cell RNA counts matrices into the reference signatures, thereby establishing a spatial mapping of cell types (4). Cell2location takes scRNA-seq derived cell type reference signatures and spatial transcriptomics data as input (2, 3). From left to right: Single-cell RNA-seq and spatial transcriptomics profiles are generated from the same tissue (1). Overview of the spatial mapping approach and the workflow enabled by cell2location. The cell2location software comes with a suite of downstream analysis tools, including the identification of groups of cell types with similar spatial locations. 
For full details and a comparison to existing approaches see our preprint (coming soon). 
Finally, (5) cell2location is computationally efficient, owing to variational approximate inference and GPU acceleration. Using these reference signatures, cell2location decomposes mRNA counts in spatial transcriptomic data, thereby estimating the relative and absolute abundance of each cell type at each spatial location (Fig 1).Ĭell2location is implemented as an interpretable hierarchical Bayesian model, (1) providing principled means to account for model uncertainty (2) accounting for linear dependencies in cell type abundances, (3) modelling differences in measurement sensitivity across technologies, and (4) accounting for unexplained/residual variation by employing a flexible count-based error model. Cell2location implements this estimation step based on Negative Binomial regression, which allows to robustly combine data across technologies and batches. 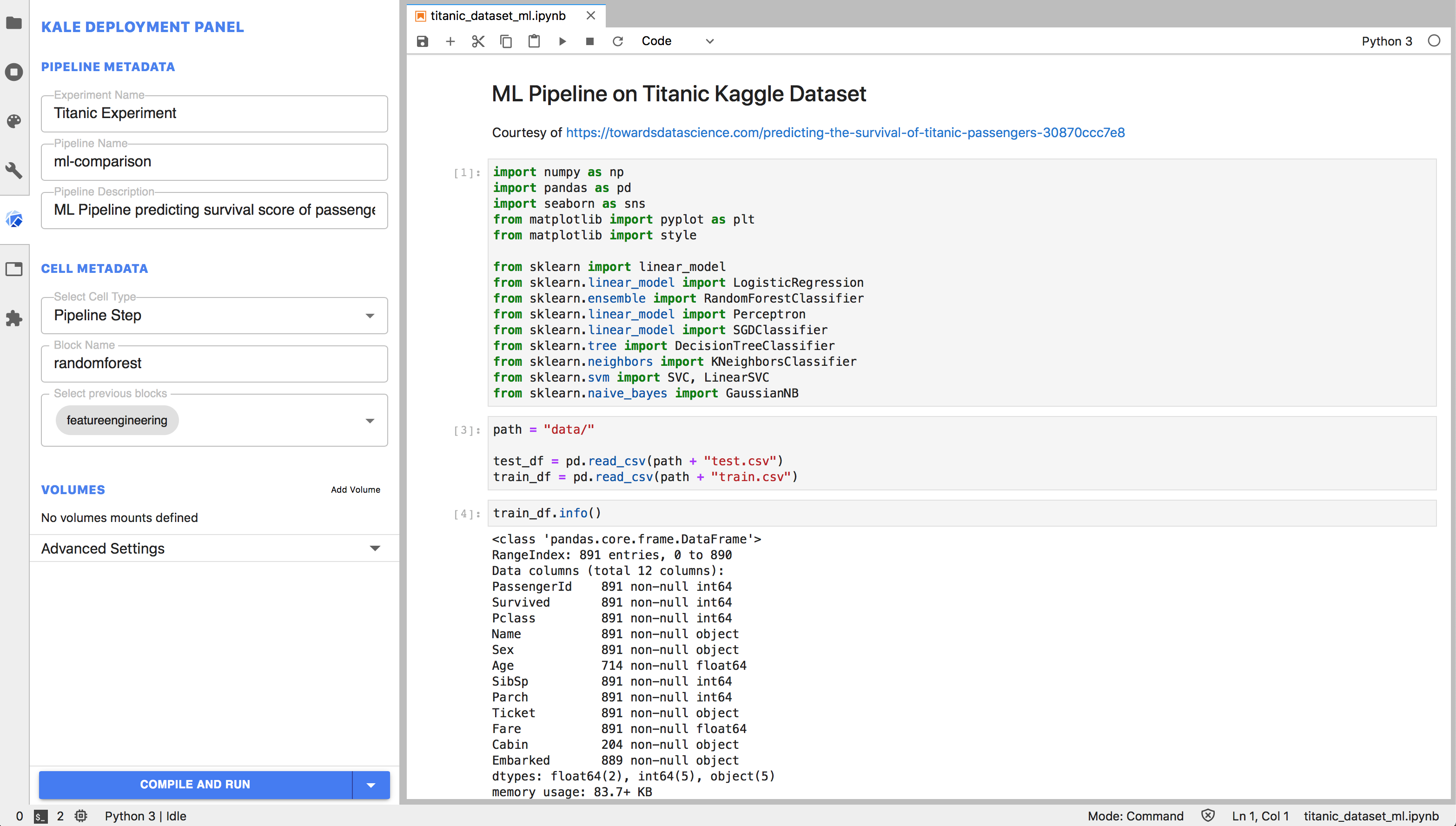
Cell2location leverages reference cell type signatures that are estimated from scRNA-seq profiles, for example as obtained using conventional clustering to identify cell types and subpopulations followed by estimation of average cluster gene expression profiles. 
Comprehensive mapping of tissue cell architecture via integrated single cell and spatial transcriptomics (cell2location model)Ĭell2location maps the spatial distribution of cell types by integrating single-cell RNA-seq (scRNA-seq) and multi-cell spatial transcriptomic data from a given tissue (Fig 1). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
